CURRICULUM VITAE

Yan Liang
Contact information
Nongshenghuan C637
Zijingang Campus, Zhejiang University
Hangzhou, 310058
China
Email: yanliang at zju.edu.cn
Phone:86-0571-88982572
Lab website: http://person.zju.edu.cn/en/yanliang
Research experience
2015-present, Professor, Zhejiang University, China
2011-2015, Postdoc fellow, University of Missouri, USA
2007-2011, Research assistant, Institute of Genetics and Developmental Biology, CAS, China
2001-2007, Ph.D., University of Vermont, USA
Research work
Molecular recognition of plant-microbe interaction
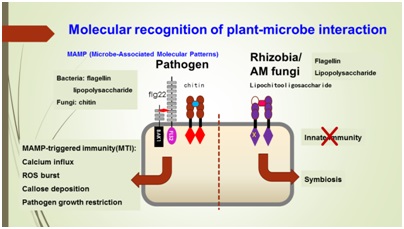
Through co-evolution with microbes, plants have evolved two opposite relationship with them. Some microbes are recognized as friends, such as rhizobia who form symbiotic relationship with legumes, whereas most of them are recognized as foes, such as pathogenic microbes. One central question in plant-microbe interactions is how plants recognize the signaling molecules from microbes to decide whether they are friends or foes. Our lab will focus on studying the recognition mechanism to some important microbial signaling molecules, including chitooligosaccharides and lipopolysaccharides.
Project I: Chitooligosaccharide (COs) and Lipochitooligosaccharide (LCO) recognition
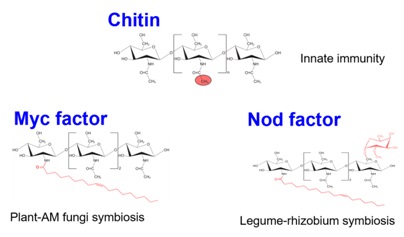
Chitin is the second most abundant polysaccharide in nature, found in crustacean shells, insect exoskeletons and fungal cell walls, and it is a β1-4 linked polymer of N-acetyl-D-glucosamine (GlcNAc). In this unmodified form, chitooligosaccharides [degree of polymerization (dp)=6-8)] are strong inducers of plant innate immunity. In contrast, when these chitooligosaccharides are acylated (so called lipochitooligosaccharides, LCO) and further modified they can act as Nod factors, the key signaling molecules that play an important role in the initiation of the legume-rhizobium symbiosis. In a similar form, these molecules can also act as Myc factors, the key signaling molecules involved in the arbuscular mycorrhiza (AM) symbiosis. The structural similarity between Nod/Myc factors and chitin raises the question how these similar molecules could induce apparently opposite responses in plants.
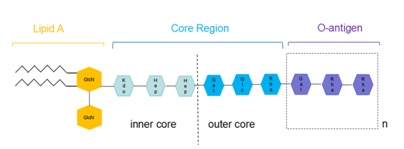
Project II: Lipopolysaccharides (LPS) recognition
Lipopolysaccharides (endotoxin) is abundant in the outer cell envelope of Gram-negative bacteria. In pathogenic bacteria, LPS acts as a MAMP that induces plant innate immune responses in various plant species, whereas rhizobial LPS might have been modified to adapt to the establishment of symbiotic relationship with legume species. Our lab will try to understand how plants recognize pathogenic LPS and how legumes have evolved to recognize rhizobial LPS.
1. Liao D, Cao Y, Sun X, Espinoza C, Nguyen CT, Liang Y, Stacey G. 2017. Arabidopsis E3 ubiquitin ligase PLANT U-BOX13 (PUB13) regulates chitin receptor LYSIN MOTIF RECEPTOR KINASE5 (LYK5) protein abundance. New Phytol. 214:1646-1656
2. Espinoza C, Liang Y, Stacey G. 2017. Chitin receptor CERK1 links salt stress and chitin-triggered innate immunity in Arabidopsis. Plant J 89(5): 984-995.
3. Stacey MG, Cahoon RE, Nguyen HT, Cui Y, Sato S, Nguyen CT, Phoka N, Clark KM, Liang Y, Forrester J, et al. 2016. Identification of Homogentisate Dioxygenase as a Target for Vitamin E Biofortification in Oilseeds. Plant Physiol 172(3): 1506-1518.
4. Xie Q, Liang Y, Zhang J, Zheng H, Dong G, Qian Q, Zuo J. 2016. Involvement of a Putative Bipartite Transit Peptide in Targeting Rice Pheophorbide a Oxygenase into Chloroplasts for Chlorophyll Degradation during Leaf Senescence. J Genet Genomics 43(3): 145-154.
5. Yang X, Nian J, Xie Q, Feng J, Zhang F, Jing H, Zhang J, Dong G, Liang Y, Peng J, et al. 2016. Rice Ferredoxin-Dependent Glutamate Synthase Regulates Nitrogen-Carbon Metabolomes and Is Genetically Differentiated between japonica and indica Subspecies. Mol Plant 9(11): 1520-1534.
6. Zhao W, Guan C, Feng J, Liang Y, Zhan N, Zuo J, Ren B. 2016. The Arabidopsis CROWDED NUCLEI genes regulate seed germination by modulating degradation of ABI5 protein. J Integr Plant Biol 58(7): 669-678.
7. Choi, Jeongmin; Tanaka, Kiwamu; Liang, Yan; Cao, Yangrong; Lee, Sang Yeol; Stacey, Gary, 2014, Extracellular ATP, a danger signal, is recognized by DORN1 in Arabidopsis, Biochemical Journal, 463, pp 429-437
8. Cao, Yangrong#; Liang, Yan#; Tanaka, Kiwamu; Nguyen, Cuong T.; Jedrzejczak, Robert P.; Joachimiak, Andrzej; Stacey, Gary, 2014, The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1, eLife, 3(e03766), (# These authors contributed equally)
9. Liang, Yan; Toth, Katalin; Cao, Yangrong; Tanaka, Kiwamu; Espinoza, Catherine; Stacey, Gary, 2014, Lipochitooligosaccharide recognition: an ancient story, New Phytologist, 204(2), pp 289-296
10. Guan, Chunmei; Wang, Xingchun; Feng, Jian; Hong, Sulei; Liang, Yan; Ren, Bo; Zuo, Jianru, 2014, Cytokinin Antagonizes Abscisic Acid-Mediated Inhibition of Cotyledon Greening by Promoting the Degradation of ABSCISIC ACID INSENSITIVE5 Protein in Arabidopsis, Plant Physiology, 164(3), pp 1515-1526
11. Choi, Jeongmin; Tanaka, Kiwamu; Cao, Yangrong; Qi, Yue; Qiu, Jing; Liang, Yan; Lee, Sang Yeol; Stacey, Gary, 2014, Identification of a Plant Receptor for Extracellular ATP, Science, 343(6168), pp 290-294
12. Van, Zhe; Hossein, Md Shakhawat; Wang, Jun; Valdes-Lopez, Osweldo; Liang, Yan; Libault, Marc; Qiu, Lijuan; Stacey, Gary, 2013, miR172 Regulates Soybean Nodulation, Molecular Plant-Microbe Interactions, 26(12), pp 1371-1377
13. Liang, Yan; Cao, Yangrong; Tanaka, Kiwamu; Thibivilliers, Sandra; Wan, Jinrong; Choi, Jeongmin; Kang, Chang Ho; Qiu, Jing; Stacey, Gary, 2013, Nonlegumes Respond to Rhizobial Nod Factors by Suppressing the Innate Immune Response, Science, 341(6152), pp 1384-1387
14. Tanaka, Kiwamu; Nguyen, Cuong; Liang, Yan; Cao, Yangrong; Stacey, Gary. 2013, Role of LysM receptors in chitin-triggered plant innate immunity. Plant Signaling and Behavior, 8(1): e22598
15. Mu, Jinye; Tan, Helin; Hong, Sulei; Liang, Yan; Zuo, Jianru, 2013, Arabidopsis Transcription Factor Genes NF-YA1, 5, 6, and 9 Play Redundant Roles in Male Gametogenesis, Embryogenesis, and Seed Development, Molecular Plant, 6(1), pp 188-201
16. Liang, Yan*; Wang, Xingchun; Hong, Sulei; Li, Yansha; Zuo, Jianru, 2012, Deletion of the Initial 45 Residues of ARR18 Induces Cytokinin Response in Arabidopsis, Journal of Genetics and Genomics, 39(1), pp 37-46 (*Corresponding author)
17. Zhang, Jian; Teng, Chong; Liang, Yan*, 2011, Programmed cell death may act as a surveillance mechanism to safeguard male gametophyte development in Arabidopsis, Protein & Cell, 2(10), pp 837-844 (*Corresponding author)
18. Yendrek, Craig R.; Lee, Yi-Ching; Morris, Viktoriya; Liang, Yan; Pislariu, Catalina I.; Burkart, Graham; Meckfessel, Matthew H.; Salehin, Mohammad; Kessler, Hilary; Wessler, Heath; Lloyd, Melanie; Lutton, Heather; Teillet, Alice; Sherrier, D. Janine; Journet, Etienne-Pascal; Harris, Jeanne M.; Dickstein, Rebecca, 2010, A putative transporter is essential for integrating nutrient and hormone signaling with lateral root growth and nodule development in Medicago truncatula, Plant Journal, 62(1), pp 100-112
19. Deng, Yan; Dong, Haili; Mu, Jinye; Ren, Bo; Zheng, Binglian; Ji, Zhendong; Yang, Wei-Cai; Liang, Yan; Zuo, Jianru, 2010, Arabidopsis Histidine Kinase CKI1 Acts Upstream of HISTIDINE PHOSPHOTRANSFER PROTEINS to Regulate Female Gametophyte Development and Vegetative Growth, Plant Cell, 22(4), pp 1232-1248
20. Chen, Ruiqiang*; Sun, Shulan*; Wang, Chun*; Li, Yansha*; Liang, Yan*; An, Fengying; Li, Chao; Dong, Haili; Yang, Xiaohui; Zhang, Jian; Zuo, Jianru, 2009, The Arabidopsis PARAQUAT RESISTANT2 gene encodes an S-nitrosoglutathione reductase that is a key regulator of cell death, Cell Research, 19(12), pp 1377-1387 (# These authors contributed equally)
21. Ren, Bo; Liang, Yan; Deng, Yan; Chen, Qingguo; Zhang, Jian; Yang, Xiaohui; Zuo, Jianru, 2009, Genome-wide comparative analysis of type-A Arabidopsis response regulator genes by overexpression studies reveals their diverse roles and regulatory mechanisms in cytokinin signaling, Cell Research, 19(10), pp 1178-1190
22. Ding, Yiliang; Kalo, Peter; Yendrek, Craig; Sun, Jongho; Liang, Yan; Marsh, John F.; Harris, Jeanne M.; Oldroyd, Giles E. D., 2008, Abscisic Acid Coordinates Nod Factor and Cytokinin Signaling during the Regulation of Nodulation in Medicago truncatula, Plant Cell, 20(10), pp 2681-2695
23. Mu, Jinye; Tan, Helin; Zheng, Qi; Fu, Fuyou; Liang, Yan; Zhang, Jian; Yang, Xiaohui; Wang, Tai; Chong, Kang; Wang, Xiu-Jie; Zuo, Jianru, 2008, LEAFY COTYLEDON1 is a key regulator of fatty acid biosynthesis in Arabidopsis, Plant Physiology, 148(2), pp 1042-1054
24. Liang Yan; Zuo JianRu; Yang WeiCai, 2007, Advances in Arabidopsis research in China from 2006 to 2007, Chinese Science Bulletin, 52(13), pp 1729-1733
25. Liang, Yan; Mitchell, David M.; Harris, Jeanne M., 2007, Abscisic acid rescues the root meristem defects of the Medicago truncatula latd mutant, Developmental Biology, 304(1), pp 297-307
26. Liang, Y; Harris, JM, 2005, Response of root branching to abscisic acid is correlated with nodule formation both in legumes and nonlegumes, American Journal of Botany, 92(10), pp 1675-1683
27. Bright, LJ; Liang, Y; Mitchell, DM; Harris, JM, 2005, The LATD gene of Medicago truncatula is required for both nodule and root development, Molecular Plant-Microbe Interactions, 18(6), pp 521-532




